Multiscale modelling of erythrocytes and mitochondrial membranes
Combining glycoproteomics, lipidomics, proteomics, structural biology, structural mass spectrometry, and integrative AI models to create testable cell membrane models.
Leads:
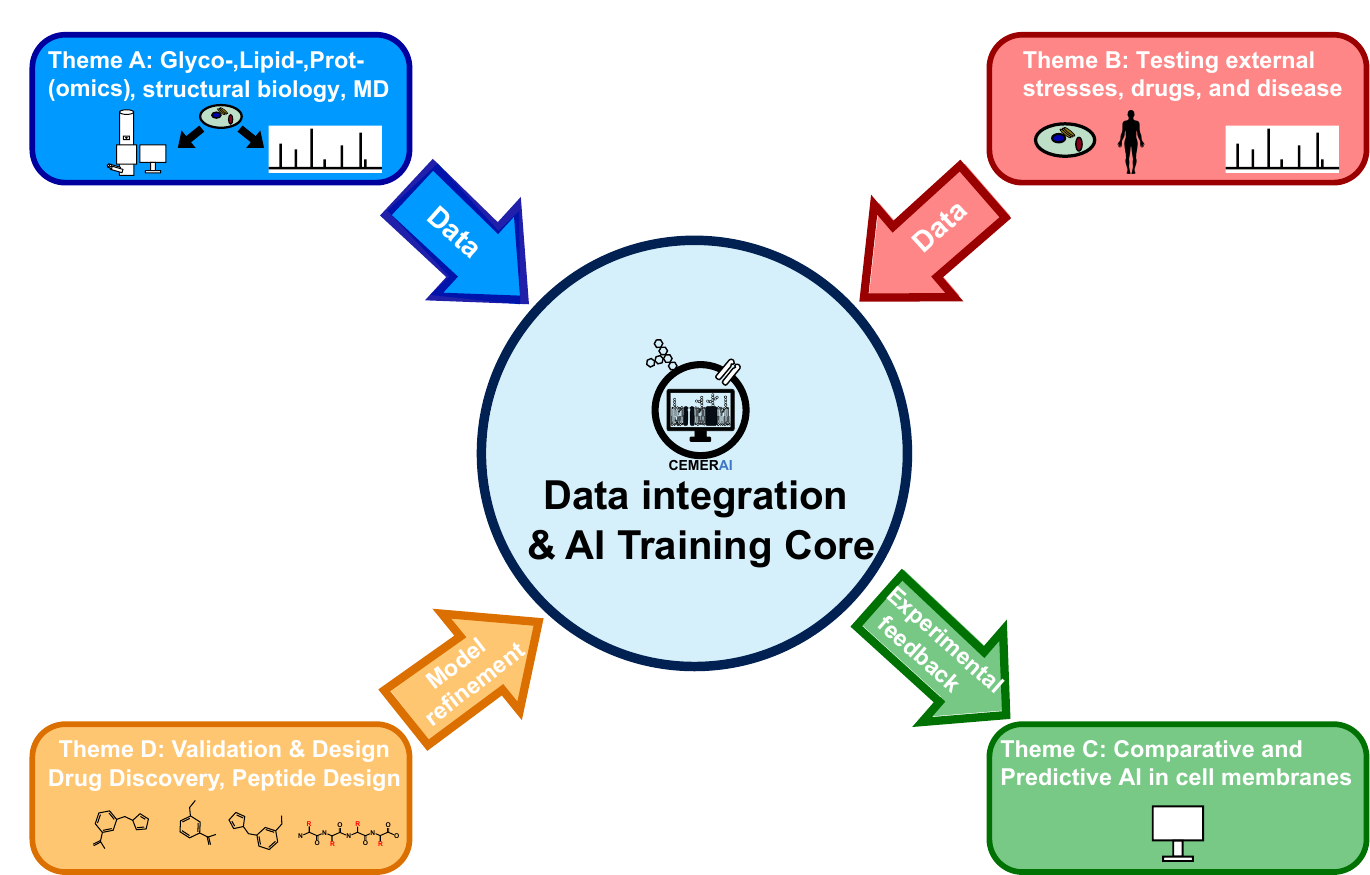
CEMERAI is designed around four interconnected scientific themes that connect data generation, modelling, prediction, and validation.
The centre integrates glycoproteomics, proteomics, lipidomics, structural biology, mass spectrometry, molecular dynamics, and AI to move from descriptive molecular catalogues toward predictive, multiscale membrane models.
| Strategic Goal | Key Scientific Questions | Core Experimental Platforms | AI/Computational Components | Expected Outputs / Impact |
|---|---|---|---|---|
| 1. Multiscale modelling of mitochondria and red blood cells | How do lipids, proteins, and glycans organize to maintain cell membrane integrity? | glycoproteomics, lipidomics, crosslinking and structural MS, cryo-EM/-ET, MD simulations | Integrative AI for multimodal coordination, ML-based feature extraction from Cryo-ET; generative AI | Predictive atomistic-mesoscale RBC/mitochondrial cell membrane model |
| 2. Modelling drug and disease-related membrane variations | How do cell membranes re-model with disease or stress or response to drugs? How do drugs enter membranes? | glycoproteomics, lipidomics, crosslinking and structural MS, cryo-EM/-ET, MD simulations, cell line and animal models | Predictive AI for dynamic cell-membrane remodelling; mapping altered lipid composition to biophysical profiles; causal modelling linking omics and phenotype | Mechanistic understanding of membrane changes in pathology; biomarkers for disease progression |
| 3. Comparative and predictive modelling of diverse cell membranes | What are the conserved and divergent structural principles across cell types and mitochondria isolated from different cell types? | Single-cell lipidomics, glycoproteomics, native MS, Cell Atlas imaging | Integrative AI for cross-cell comparison; transfer learning between models | Cross-cellular membrane atlas; digital reference for drug response and tissue-specific therapeutics |
| 4. Use of synthetic protein design for diagnostics and therapeutics | Development of advanced data-driven tools for diagnostics and treatment | Synthetic biology, high-throughput binder design | Generative AI for peptide/protein binder design | New therapeutic and diagnostic molecules; open-access AI models for the life science community |
Combining glycoproteomics, lipidomics, proteomics, structural biology, structural mass spectrometry, and integrative AI models to create testable cell membrane models.
Leads:
Identifying plasma membrane and mitochondrial remodelling events during pathological, drug-induced, or stress-induced states.
Leads:
Building cross-cell-type models to capture conserved and specialised principles of membrane organisation for diagnostics and rational therapeutic design.
Leads:
Using AI tools to develop synthetic binders against key membrane proteins, including glycoproteins with high specificity.
Leads:
Here we will develop generative and predictive AI models to improve our understanding of drug transport into cells.
Leads: